Shadow Contact Map
A contact map is an important component of a structure-based model. It is a symmetric matrix that encodes the tertiary structure of the molecule by defining the interactions that stabilize the native state. A common method of choosing these interactions is to choose a cut-off distance and define all atoms within the cut-off distance in the native state as the stabilizing interactions. The cut-off distance needs to be long enough to include all relevant short range interactions, for proteins ~6Å should be sufficient. With a cut-off definition for contacts, in order to include the contacts between 5-6Å, several "unphysical" contacts will be introduced. The unphysical contacts being those that are acting through another atom. To avoid this situation we introduce the Shadow contact map (SCM).
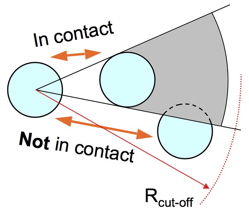
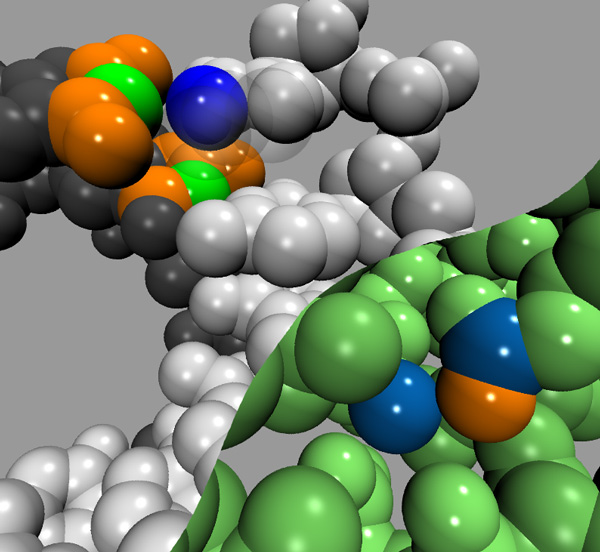
The SCM algorithm is described in the above figure. It is a cut-off contact map with a screening term S introduced. All atoms are given a radius S. To determine the atoms contacting atom i, all atoms within a cutoff distance C of atom i are considered possible contacts. Those contacts which have an intervening atom are considered screened out and are discarded. The algorithm is simply stated: a light source is located at the center of atom i, any atoms within C which have no shadow cast upon them are considered contacts. One can change S to scale the amount of shadowing. S=1Å works well with proteins. The figure below highlights SCM in action. The upper figure shows an RNA double helix. With a cut-off of 4Å the blue atom is in contact with the orange and green atoms. With a shadowing radius of 1Å, only the green atoms are in contact with the blue. The lower figure shows the interior of chymotrypsin inhibitor 2 (PDB code 2CI2). The blue atoms are contacting with a 4Å cut-off map, but with S>0.5Å the orange atom shadows and throws out the contact.
Details are described in the following reference:
Noel JK, Whitford PC, Onuchic JN. (2012) "The shadow map: A general contact definition for capturing the dynamics of biomolecular folding and function" J. Phys. Chem. B DOI:10.1021/jp300852d
Shadow (SCM.jar) software
SCM.jar is distributed as part of the SMOG 2 software bundle, available here. See SMOG 2 user manual for usage details.HOW SCM WORKS:
- Determines which atoms are in contact using the following algorithm:
- Checks if two atoms are within cutoff (-c) angstroms.
- Checks if two atoms are greater than the specified number of residues apart or in separate chains.
- If so then checks to see if any atoms are between the two. This is determined by putting a light bulb at the center of each of the atoms and seeing if any of the other atoms cast a shadow on the potential contacting atom. If so then the contact is thrown out.
- The size of the atoms is determined by size (-s).
- A
potentially shadowing atom is given a size of 0.5 angstrom when it is
bonded to the shadowee. This is because the C-C bond length
is 1.5
angstrom and bonded neighbors should not shadow.
This resource is provided by the Center for Theoretical Biological Physics.
Please direct questions and comments to info@smog-server.org.
Page created and maintained by Jeff Noel and Paul Whitford
